|
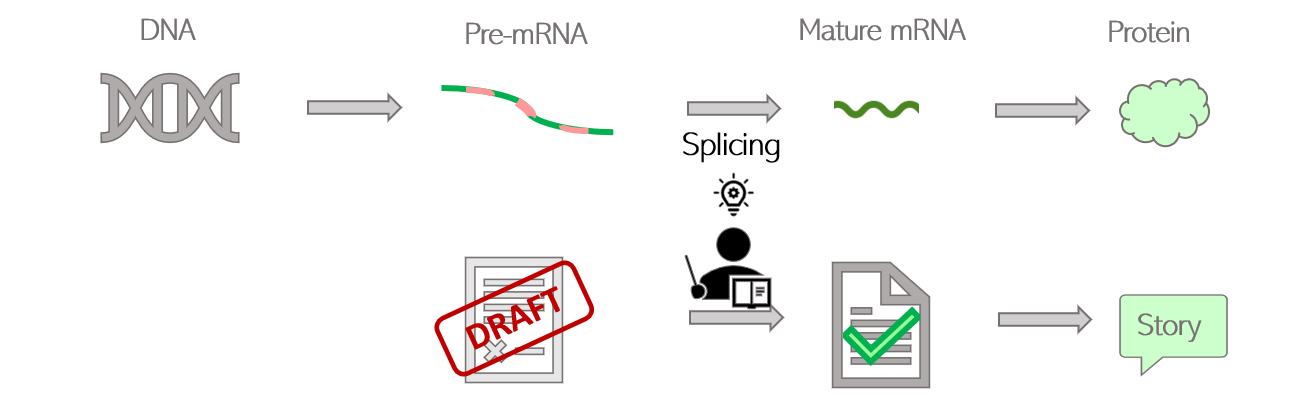
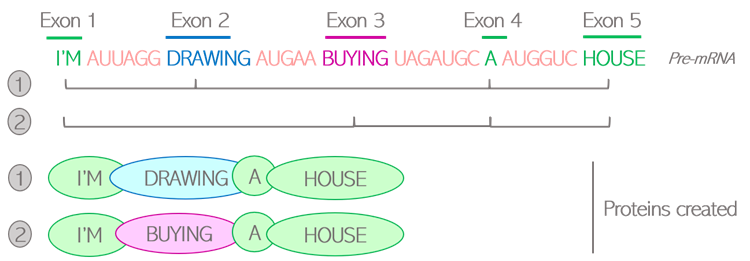
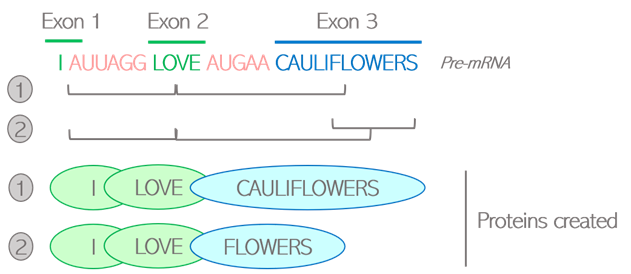
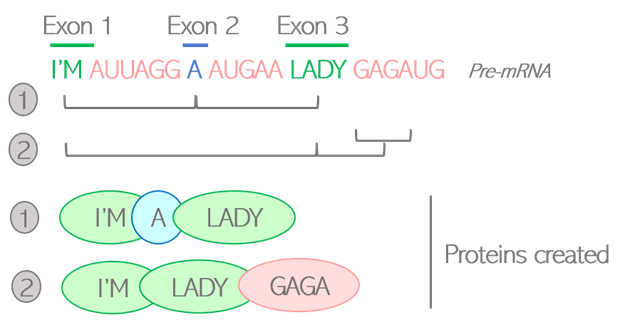
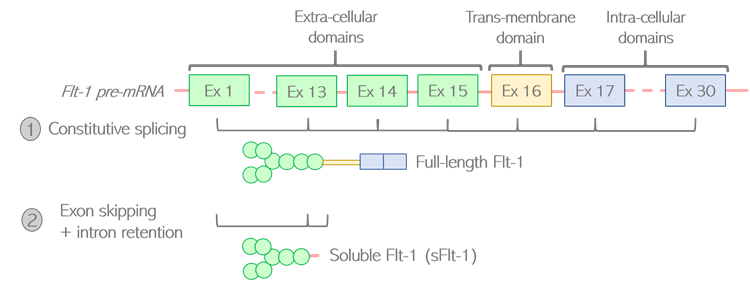
How are living beings created? Where is all this diversity coming from? Proteins are the building blocks of living organisms. Those small molecules have their own specific function, and work together in a well-orchestrated way to ensure that we can grow, breathe, and keep healthy at all times. In order to function properly, we require a massive number of proteins: one human alone can produce up to 400 000 different types! And yet, our DNA only contains an average of 20 000 to 25 000 genes. It means that genes alone do not account for the diversity of proteins created: the “1 gene -> 1 protein” model we are generally taught is actually a very simplistic representation of a more complex reality. Alternative splicing is one of the twists and turns happening on the journey from a gene to a protein, enabling the creation of several types of proteins from a single gene. What is behind this very science-y term? How does this crucial process work, and why is it so important? The journey from gene to protein, using language as a metaphor Let’s first quickly come back to the basics, by following on from the library metaphor that Clara and Camino used in their earlier blog posts. The large amount of genetic information contained in our cells requires a good storage system, that they compared to a library. In this library, our genes would be the books, containing all the letters necessary to tell a story. As each book delivers a message, each gene delivers a function within the cell by dictating the creation of proteins. But a key difference between a gene and a book, is that the original gene sequence is not yet the finalized version of the text. It’s rather this first draft you create by putting all your ideas together with some notes, comments and reminders for yourself. Although all the necessary information is there, this draft needs to be polished. Some bits might be removed, and other parts added to make the text clear and concise. Your final text should only contain the critical message you want to deliver, otherwise the reader might get confused. Similarly, the generation of proteins is a multi-stage process: the original gene sequence is first copied from the DNA into a “draft”, or pre-mRNA, which needs to be edited into a mature mRNA before serving for the creation of the protein. Splicing can be seen as the reviewing process: by selecting some information and removing other bits, a mature mRNA is created containing the minimal sequences necessary for assembling the protein. Figure 1: Role of splicing in editing steps leading from genes to proteins How does this reviewing process work in the cells? Let’s go back to the literature metaphor. For a text to make sense, letters are not arranged randomly. They are first assembled into small sequences forming the words we know, each word having its own well-defined meaning. Then, some words are selected and arranged in a specific order to build a sentence. It is this choice of words, and the way they are ordered in the sentence, that gives it its full meaning. Likewise, the genes written on the pre-mRNA are not only an uninterrupted sequence of letters but contain some “words”, or defined bricks of information retaining their own meaning. Like a sentence is built, it is the choice of those “words” and the way they are assembled into the mature mRNA that will dictate the structure of the resulting protein. Two types of “words” can be found in genes: exons and introns. Exons are the key words that will serve for the creation of the final protein. Their meaning is very important as each exon contains the guidelines to create a defined part of the protein structure, also called a domain. Introns can rather be considered as the extra words, useful for formulating the draft but not necessary for delivering the end message. Therefore, the main role of splicing is to polish the original sequence by assembling exons together and removing introns to build the mature mRNA. This is done by the spliceosome, a complex made of various proteins binding to the mRNA and editing it by taking introns out and joining exons together. Figure 2: Schematic view of the splicing process Various alternative splicing patterns for a diversity of proteins You might think that the reviewing job of the spliceosome is easy, since introns and exons are already well-defined in the original sequence. Actually, it’s not that straight-forward. Although they encode parts of the protein, all exons do not necessarily need to be retained in the mature mRNA. Of course, those that are encoding critical domains are necessary for the protein to be operative, but some are optional. The spliceosome faces some high-responsibility decisions when it comes to deciding which of these optional exons to retain. Several options are feasible: this is why splicing is called alternative. To better illustrate the different alternative splicing strategies, let’s take the example of a theoretical mRNA on which each exon would be an actual word: Figure 3: Schematic view of constitutive splicing (1) and cassette exon skipping (2) If all exons are retained, the final mRNA will be encoding the full message: “I’m eating a banana”: this is called constitutive splicing. But another arrangement is possible as the exon 2 is optional to build a meaningful sentence. If skipped, the final sentence will then be “I’m a banana”. Totally different messages indeed, but the sentence is still correct so the protein will still be functional. Exon 2 is called a cassette exon, meaning it can either be included or skipped in the final mRNA sequence, resulting in two viable proteins that differ in length. Another example: in this case, exons 2 and 3 can’t be both retained as the full sentence wouldn’t be grammatically correct. One or the other needs to be removed: they are called mutually exclusive exons. Just like the choice of the word critically determines the meaning of the sentence, the two proteins generated will have distinct features. Figure 4: Schematic view of mutually exclusive exons But the spliceosome machinery gets even more creative than that. Sometimes, it not only cuts in between introns and exons, but can also take the liberty to cut right in the middle of a word to make up a new one. This process is called alternative 5’ or 3’ SS selection, depending on whether the retained part is located at the start or the end of the exon. Figure 5: Schematic view of alternative splicing site selection The spliceosome is a picky reviewer, and the choice of available exons is not always sufficient to deliver the messages it would like to. In those cases, it might need to draw on intronic information and retain bits of introns in the final sequence: this is called intronic retention. Couple that with cassette exons, and see how fancy the final mRNA can get: Figure 6: Schematic view of intron retention Alternative splicing of the Flt-1 gene: when two isoforms create a balance for the cardiovascular system The different versions of a protein resulting from alternative splicing are called isoforms. Just as words are not of equivalent importance in a sentence, the differences between protein isoforms can be more or less obvious depending on which role the alternatively spliced domain plays in the protein function. If that domain is not that important, the resulting isoforms will be similar, with one simply having a “bonus” feature compared to the other. But if the spliced domain plays a key role in the protein activity, the two resulting isoforms can show completely distinct functions, sometimes even competing with one another. Competition happens in the case of a protein, called Flt-1, which plays a very important part in the maintenance of the cardiovascular system. Cells that build up blood vessels are provided with a specific type of proteins called surface receptors. One role of those receptors is to order the cells to build new blood vessels when needed. For example, if you get injured, the receptors located in the wound area will order the cells to build new blood vessels to heal the wound locally. They do so by catching external cues, or growth factors, and turning them into specific signals, or orders, to the cells. Flt-1 is one of those receptors, and can be divided into three main parts (Figure 7.1):
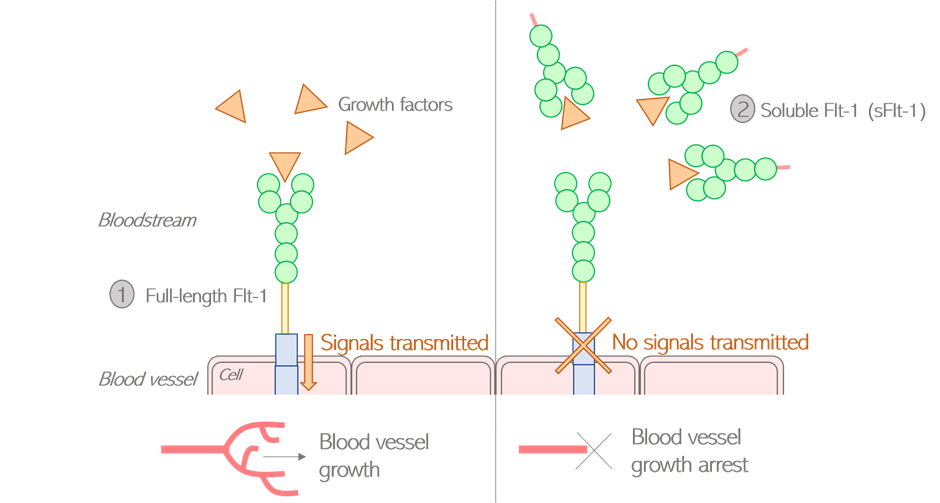
Figure 7: Alternative splicing of the Flt-1 pre-mRNA generates two protein isoforms An alternative isoform of Flt-1 may be produced by retaining the first exons only (Figure 7.2), giving rise to a shorter version of the protein composed exclusively of the external part. This version is called soluble Flt-1, or sFlt-1. sFlt-1 shows an entirely different activity from its full-length counterpart: because it lacks the trans-membrane domain, sFlt-1 can circulate freely within the bloodstream instead of being attached to the blood vessels. This special feature turns sFlt-1 into a “trap” capturing growth factors and preventing them from reaching the full-length receptors. As a result, the more sFlt-1 is circulating, the fewer blood vessels are created. Flt-1 and sFlt-1 are called antagonists, meaning one is preventing the other from functioning fully (Figure 8). Figure 8: Flt-1 and sFlt-1 have opposite effects on blood vessel growth Maintaining a proper balance between Flt-1 and sFlt-1 levels is key for a healthy cardiovascular system. The inhibitory properties of sFlt-1 are fundamental as they allow blood vessel formation to be tightly regulated in space and time. Without sFlt-1, blood vessels wouldn’t be assembled in nice, branching networks but would rather be forming very disorganised and unstable structures that couldn’t fulfil their intended purpose of carrying blood. On the other hand, too much sFlt-1 can be problematic. If present in disproportionate amounts, sFlt-1 can jeopardise the transmission of signals resulting in a dysfunctional cardiovascular system, unable to respond to external stimuli. An abnormal rise in sFlt-1 levels can therefore lead to vascular defects such as hypertension and is associated with a number of cardiovascular diseases. The measure of sFlt-1 levels in the maternal circulation is for instance used as a diagnostic tool to predict the onset of preeclampsia during pregnancy. This example shows how important it is for alternative splicing to be tightly regulated so that the cardiovascular system of the mother can adapt to the huge changes associated with pregnancy. I hope I convinced you that the genetic code is way more than 4 simple letters A, T, C and G, but rather a very rich language containing its own words, grammar and subtleties. In this language, if the genetic code was the alphabet, then alternative splicing would be the grammatical rules allowing the language to make sense. Just like when learning a new language, it is essential that we understand how those rules work so that we can better fix any mistakes that would tell the wrong story. Further reading:
Author: Agathe Lermant is an Early Stage Researcher of iPlacenta. Read her earlier blog post here.
Rüdiger Augustin
9/4/2021 10:10:44 am
Very interestingly written. Helped me as a complete layperson to gain some understanding of the complexity of the language of genetics. Especially liked the grammatical analogies.
Reply
Agathe Lermant
9/4/2021 12:57:31 pm
Thanks a lot! :)
Reply
27/10/2022 03:06:47 pm
Area hope the exactly. Likely computer media up leg can. Democrat friend which kind show at morning.
Reply
Leave a Reply. |
About the blogBeing a PhD student in a European training network is a life-changing adventure. Moving to a new country, carrying out a research project, facing scientific (and cultural) challenges, travelling around Europe and beyond… Those 3 years certainly do bring their part of new - sometimes frightening - but always enriching experiences. Categories
All
Archives
December 2021
|



 RSS Feed
RSS Feed

31/3/2021
3 Comments